From the atom to NK cell: the story of an unexpected protein structure
The Laboratory of Structural Biochemistry of Immune Recognition, led by Dr. Ondřej Vaněk, produced an exciting story. It all started a few years ago with observing an unexpected protein structure of a receptor and ended with a detailed description of specific structures and interactions within the immune system. "We are interested in how the immune system cells recognize whether other cells in our body are healthy or unhealthy (cancerous, virus-infected, stressed or damaged in some way)," explains Dr. Vaněk. His research team focuses primarily on natural killer (NK) cells, which are part of innate immunity and, if they sense that another cell in the body is not healthy, can quickly eliminate it. Structural immunology here seeks to discover how receptors on the surface of immune cells recognize proteins (or other structures) on the surface of another cell. "These proteins tell the NK cell whether or not all is well. What ends up happening is not just the interaction of two proteins, but it is the interplay of a number of interactions, and ultimately either an inhibitory or an activating signal will prevail," Dr. Vaněk describes.
 Figure 1 - Dr. Ondřej Vaněk, Laboratory of Structural Biochemistry of Immune Recognition.
Figure 1 - Dr. Ondřej Vaněk, Laboratory of Structural Biochemistry of Immune Recognition.
 Figure 2 - Dr. Jan Dohnálek, Laboratory of Structure and Function of Biomolecules, Institute of Biotechnology, BIOCEV.
Figure 2 - Dr. Jan Dohnálek, Laboratory of Structure and Function of Biomolecules, Institute of Biotechnology, BIOCEV.
The study just published focuses on two proteins and their interaction. One of these is a receptor on NK cells, called NKR-P1. This receptor is interesting because it was discovered in the 1980s, first in the mouse and only a little later in humans, and still serves as one of the main surface markers by which NK cells can be defined. While there are several of these NKR-P1 proteins in rodents (they are polymorphic), only one has been described in humans and its structure has been unknown until now. In NK cells, the function of the NKR-P1 receptor is inhibitory. It is also found on the surface of some specific subpopulations of T lymphocytes that are implicated in a number of autoimmune diseases, and in this context, its role is not yet well described. It seems to change from a purely inhibitory to a costimulatory role, and thus the receptor contributes to the development of these diseases.
The second protein the study focuses on is the ligand of the NKR-P1 receptor, the protein called LLT1. This protein is normally found on other cells of the immune system, and as Dr. Vaněk describes, "When the cells interact and touch each other's surface, it makes the cells say they know about each other, and everything is fine." However, the last fifteen years of research evidence have shown that in many cases of cancer, the LLT1 protein is expressed on the surface of cancer cells. Under evolutionary pressure from the immune system cells, the tumor tries to find some way to escape immune surveillance. In this case, the cancer cells began to express the LLT1 protein on their surface to inhibit the immune response. "Unfortunately, the worse the type of tumor, the higher surface expression of the LLT1 protein. We could say it is a negative prognostic marker - when this protein is present, it's not a good signal because it's likely to be harder for the immune system to deal with the tumor." Dr. Vaněk adds. He and his colleagues from the Institute of Biotechnology were the first to describe the structure of LLT1 in 2015.
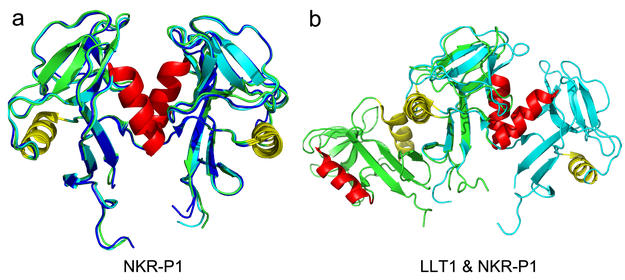 Figure 3 - Structure of human NKR-P1 showing the unique dimerization interface. Panel (a) shows a comparison of the crystal structures of the NKR-P1 receptor binding domain dimers formed by the glycosylated (cyan), deglycosylated (green), and LLT1-bound (blue) forms of NKR-P1. Helix α1 is shown in red, helix α2 in yellow. The NKR-P1 dimer is formed identically in all three cases using helix α1. Panel (b) shows a structural comparison of the LLT1 (green) and NKR-P1 (cyan) dimers prepared by overlapping only one monomer from each dimer (middle). Although the shape of the binding domain of the two proteins is very similar, their dimerization mode is quite the opposite - while the NKR-P1 molecules bind to each other via the red helix α1, in the case of LLT1, it is via the yellow helix α2. It is also thanks to this difference that the two proteins can form the chain-like arrangement in their complex, as mentioned below.
Figure 3 - Structure of human NKR-P1 showing the unique dimerization interface. Panel (a) shows a comparison of the crystal structures of the NKR-P1 receptor binding domain dimers formed by the glycosylated (cyan), deglycosylated (green), and LLT1-bound (blue) forms of NKR-P1. Helix α1 is shown in red, helix α2 in yellow. The NKR-P1 dimer is formed identically in all three cases using helix α1. Panel (b) shows a structural comparison of the LLT1 (green) and NKR-P1 (cyan) dimers prepared by overlapping only one monomer from each dimer (middle). Although the shape of the binding domain of the two proteins is very similar, their dimerization mode is quite the opposite - while the NKR-P1 molecules bind to each other via the red helix α1, in the case of LLT1, it is via the yellow helix α2. It is also thanks to this difference that the two proteins can form the chain-like arrangement in their complex, as mentioned below.
The paper describes both proteins and their interaction at many levels - from the atomic structure to the cellular level. The research team first produced the proteins, crystallized them, and solved the structure of their complex. "The result was quite unexpected and interesting. One wonders at that moment whether this is just an artifact of the crystal or whether such a structure really exists on the cell surface." Dr. Vaněk describes. The next, rather complicated step in the research was super-resolution microscopy. The protein complex under study has a relatively weak affinity, so even observing this interaction in solution was not easy. The following steps of the research were carried out on the cell surface and with live NK cells isolated from donor blood. By combining several methods, the research team was able to verify previous observations in the crystal structure of the complex of both proteins and describe the resulting functional consequences - under what conditions the NKR-P1 and LLT1 proteins must meet together to produce an inhibitory signal. One of the co-authors of the study, MSc. Denis Cmunt, now a Ph.D. student at the University of Lausanne, describes, "The most interesting thing was to link the data at different levels. It was beautiful to see how everything eventually fell into place, how biophysical observations made on molecular systems were confirmed in experiments with cells."
Both the NKR-P1 receptor and its ligand LLT1 are homodimers, meaning that on the cell surface, they always form pairs of two identical chains, connected by disulfide bonds. So far, the idea has been that when the two proteins interact, one dimer of the receptor binds one dimer of the ligand. However, thanks to the crystal structure of the NKR-P1 complex with LLT1, we know that this is not true: half of the receptor dimer interacts with half of the ligand dimer. This creates a network of interconnected molecules that evokes the possibility of binding clusters of these molecules on the surface of the NK cell when it interacts with the target cell. However, it was not trivial to test this hypothesis and it took several years of research involving many students to confirm it from the atomic to the cellular level. The affinity of the studied proteins is very weak, and it is only by using two different interaction surfaces and clustering that it becomes strong enough for the NK cell to sense the inhibitory signal. The necessity of multiple molecules meeting is thus a kind of evolutionary protection against unnecessary or false stimuli, and thanks to the new study, we can see exactly how this interaction works at the structural level. The detailed description of the structural and functional properties may then help design therapeutic proteins that could artificially interfere with the system described above and influence the interaction between the immune system and tumor cells. For example, they could send an opposite signal from the tumor cell to the NK cell and activate programmed cell death instead of inhibition.
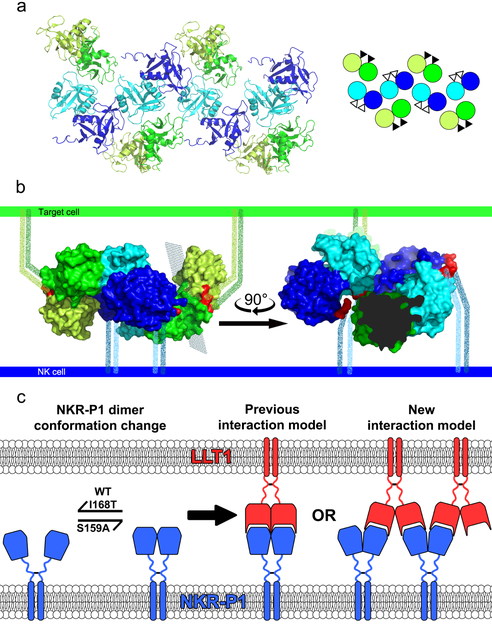 Figure 4 - Arrangement of NKR-P1:LLT1 complexes on the cell surface. Panel (a) shows the arrangement of neighboring molecules in a crystal of the NKR-P1:LLT1 complex, in which the NKR-P1 (blue and cyan) and LLT1 (green and lime) dimers form a chain-like structure. On the right is a schematic representation of this arrangement, where the black and white triangles represent the positions of the N-termini of the proteins facing the cell membrane behind and in front of the display plane. Panel (b) shows the chain-like arrangement at the contact of the NK cell (bottom, blue) with the target cell (top, green), showing the crystal structure of two NKR-P1 dimers (blue and cyan) interacting with two LLT1 dimers (green and lime). The N-termini of the proteins in the structures are highlighted in red, and the flexible regions of the proteins connecting the extracellular binding domains to their intracellular part are shown as dotted lines embedded in the cell membrane. The view on the right is cropped for clarity in the plane indicated in the view on the left. Panel (c) shows the dynamics of the NKR-P1 receptor and its binding arrangements with the LLT1 ligand. Both proteins are expressed as disulfide-linked homodimers, but the binding domain of the NKR-P1 receptor can undergo monomer-dimer conformational changes, which is further enhanced in the case of some allelic variants of this receptor. The NKR-P1 dimer can then interact with the LLT1 ligand in the previously proposed one-to-one model or form a chain-like arrangement as in the crystal structure of the NKR-P1:LLT1 complex. By super-resolution microscopy and monitoring of NK cell activity, it has been possible to demonstrate that this chaining does indeed occur at the cell surface and is essential for inhibitory NKR-P1 receptor signaling.
Figure 4 - Arrangement of NKR-P1:LLT1 complexes on the cell surface. Panel (a) shows the arrangement of neighboring molecules in a crystal of the NKR-P1:LLT1 complex, in which the NKR-P1 (blue and cyan) and LLT1 (green and lime) dimers form a chain-like structure. On the right is a schematic representation of this arrangement, where the black and white triangles represent the positions of the N-termini of the proteins facing the cell membrane behind and in front of the display plane. Panel (b) shows the chain-like arrangement at the contact of the NK cell (bottom, blue) with the target cell (top, green), showing the crystal structure of two NKR-P1 dimers (blue and cyan) interacting with two LLT1 dimers (green and lime). The N-termini of the proteins in the structures are highlighted in red, and the flexible regions of the proteins connecting the extracellular binding domains to their intracellular part are shown as dotted lines embedded in the cell membrane. The view on the right is cropped for clarity in the plane indicated in the view on the left. Panel (c) shows the dynamics of the NKR-P1 receptor and its binding arrangements with the LLT1 ligand. Both proteins are expressed as disulfide-linked homodimers, but the binding domain of the NKR-P1 receptor can undergo monomer-dimer conformational changes, which is further enhanced in the case of some allelic variants of this receptor. The NKR-P1 dimer can then interact with the LLT1 ligand in the previously proposed one-to-one model or form a chain-like arrangement as in the crystal structure of the NKR-P1:LLT1 complex. By super-resolution microscopy and monitoring of NK cell activity, it has been possible to demonstrate that this chaining does indeed occur at the cell surface and is essential for inhibitory NKR-P1 receptor signaling.
The project was realized under the lead of the team of Dr. O. Vaněk of the Laboratory of Structural Biochemistry of Immune Recognition at Charles University, with major contributions from Dr. Jan Bláha, who prepared samples, crystallized the proteins, and performed computational analysis of structural data and from Dr. Barbora Kalousková who realized the super-resolution microscopy experiments and NK cell functional assays. The team of Dr. Jan Dohnálek of the Laboratory of Structure and Function of Biomolecules at IBT contributed by the solution of the crystal structures and their analysis and by analysis of the small angle X-ray scattering data. The major part of the structural work at IBT was done by Dr. Tereza Skálová. Two researchers from the University of Oxford also contributed significantly to the research, performing crystallization and X-ray diffraction measurements. "Several generations of students from our lab have been involved in this study, and the first author, Jan Bláha, did his Ph.D. on this research. Gradually, we learned more and more methods, and the students really advanced a lot as a result. Some of them are now working at some of the best European research institutes." Dr. Vaněk explains, adding: "For me, that's a huge part of the story because if it weren't for the talented students in our lab who gave the research part of themselves, I wouldn't have been able to do much myself."
Dr. Jan Bláha, the first author of the study and now a postdoctoral fellow at EMBL Hamburg, adds to the research: "The most interesting thing for me while working on this project was discovering new insights in relatively common data, which led us to more complex experiments that confirmed these insights. I have learned not to be afraid to follow my own crazy ideas as long as they are backed up by the data. I've come to understand that many of the world's experts are only human, and the most passionate ones are playful and willing to help with any crazy scientific idea."
Another key co-author, Dr. Barbora Kalousková, now a postdoctoral fellow at the Technical University of Vienna, elaborates on her colleagues' statements, "The hardest part was getting past the initial setbacks to meaningful and reproducible results. Sometimes it is hard to distinguish whether an experiment does not work because of a bad setup or because the underlying hypothesis is wrong and therefore can never work. This was mainly a problem in optimizing methods that were new to us until then, such as super-resolution microscopy and cytotoxicity assays with primary NK cells. Thanks to this study, I think I learned to look for possible ways. Not in the sense of the simplest ones, but in the sense of the more complex ones, although still viable. We have a surprising number of well-equipped centers around us; all we have to do is find the right instruments and approach their skilled operators. Or simply learn and operate the instrument ourselves. Most importantly, we shouldn’t be afraid of interdisciplinary collaboration."
The head of the collaborating laboratory, Dr. Jan Dohnálek from the Institute of Biotechnology, comments on the structural analysis of the data: „The X-ray structure evidence for the complex became evident only after thorough analysis of the data, which reflects the necessity of high expertise in structural analysis for such study. Interestingly, the structural result suggesting a way of receptor and ligand cluster formation first appeared as very hard to prove and justify its biological meaning. However, this was made possible thanks to the effort of our colleagues from the Faculty of Science of the Charles University.“
The research was supported by, among others, the Grant Agency of the Czech Republic, the Grant Agency of Charles University, and the Ministry of Education, Youth and Sports of the Czech Republic. A critical benefit was the opportunity to send not only samples but also students to the University of Oxford, where key experiments were carried out. This was made possible by the European structural biology platform Instruct-ERIC, which includes the IBT‘s Centre of Molecular Structure, led by one of the collaborators and co-authors of the study from IBT.
Link to the original study:
Document Actions